 The fact that the green reads are pointing off the edges of the reference and we know they are 250bp molecules, leads to one structural solution as circular molecules. The top track is a depiction of a linear plasmid assembly and the reads with in that assembly that are evidence of the circular state of the molecule. The middle plasmid is a depiction of the circularized read pairs from the Top track. The vector on the right is what the RNase A sequencing and assembly produced. The Broad Institute (the authors of IGV:Integrated Genome Viewer used in Figure 1), have a great tutorial that covers read pairing with Illumina and SOLiD sequencing.įigure 5 Illustrates Pfizer’s disclosed vector map (EMA documentation below) on the left. Detection of large deletions with insert size deviations after read mapping. The tighter your size distribution of molecules, the easier it is to detect deviations from that distribution.įigure 4. We were very careful to make these very tight insert size libraries so we have an expected distance between the forward and reverse sequencing reads. Figure 2 is the DNA length of the RNase A treated libraries. Let us expand on this point as it a very important aspect of next generation sequencing.  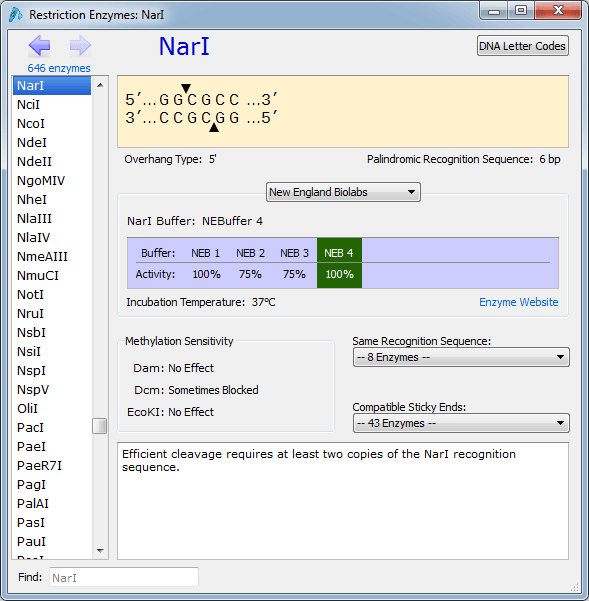
Since these sequencing libraries were known to be 250 base pairs in size, we can look for reads that map ~7,700 bases apart and point across the circularization point. This enabled a deeper sampling of the DNA based molecules without the mRNA sequence obscuring and suppressing the depth of coverage for the plasmid vector.Īfter assembling and mapping these paired reads back to the reference genome we can look for reads that point across the part of the reference that should be circular. RNase A digests RNA into small pieces that can be eliminated with purification. To address this problem, we RNase A treated the sample and sequenced the remaining DNA. This made it difficult to assess the % of the plasmid DNA that was linear versus circular DNA. The RNA-Seq kit employed favored sequencing RNA and under estimated the DNA contamination levels in the vaccine. 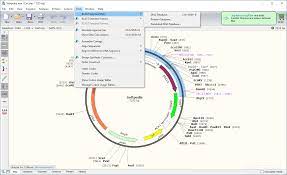
Previous work sequencing the Pfizer bivalent vaccines utilized a method that sequenced both the RNA and DNA in the sample. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
